1. Navigate to BaseSpace Import
To import samples from BaseSpace, first click “Upload” on the left navigation bar. Click “Illumina BaseSpace”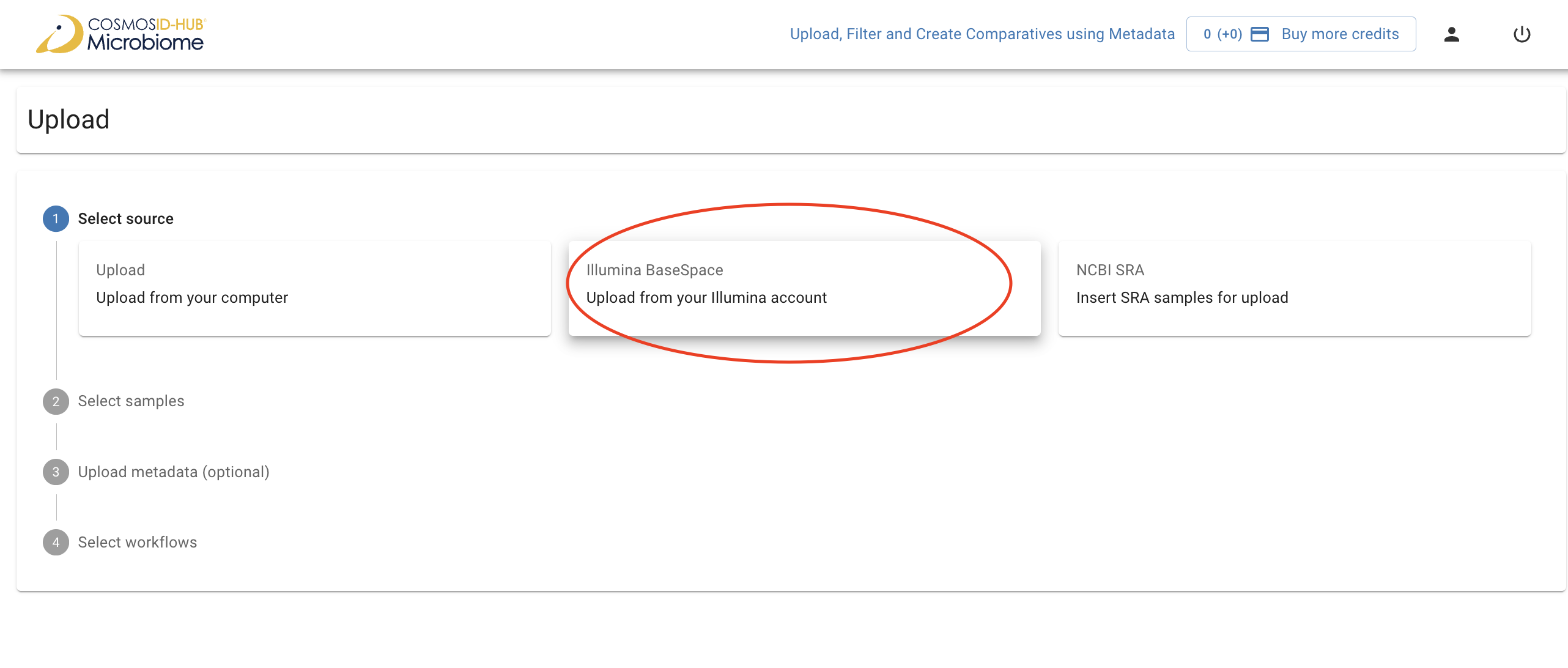
2. Log Into BaseSpace
A window will open that allows you to log in to your BaseSpace account. Enter your Illumina BaseSpace account email address and password.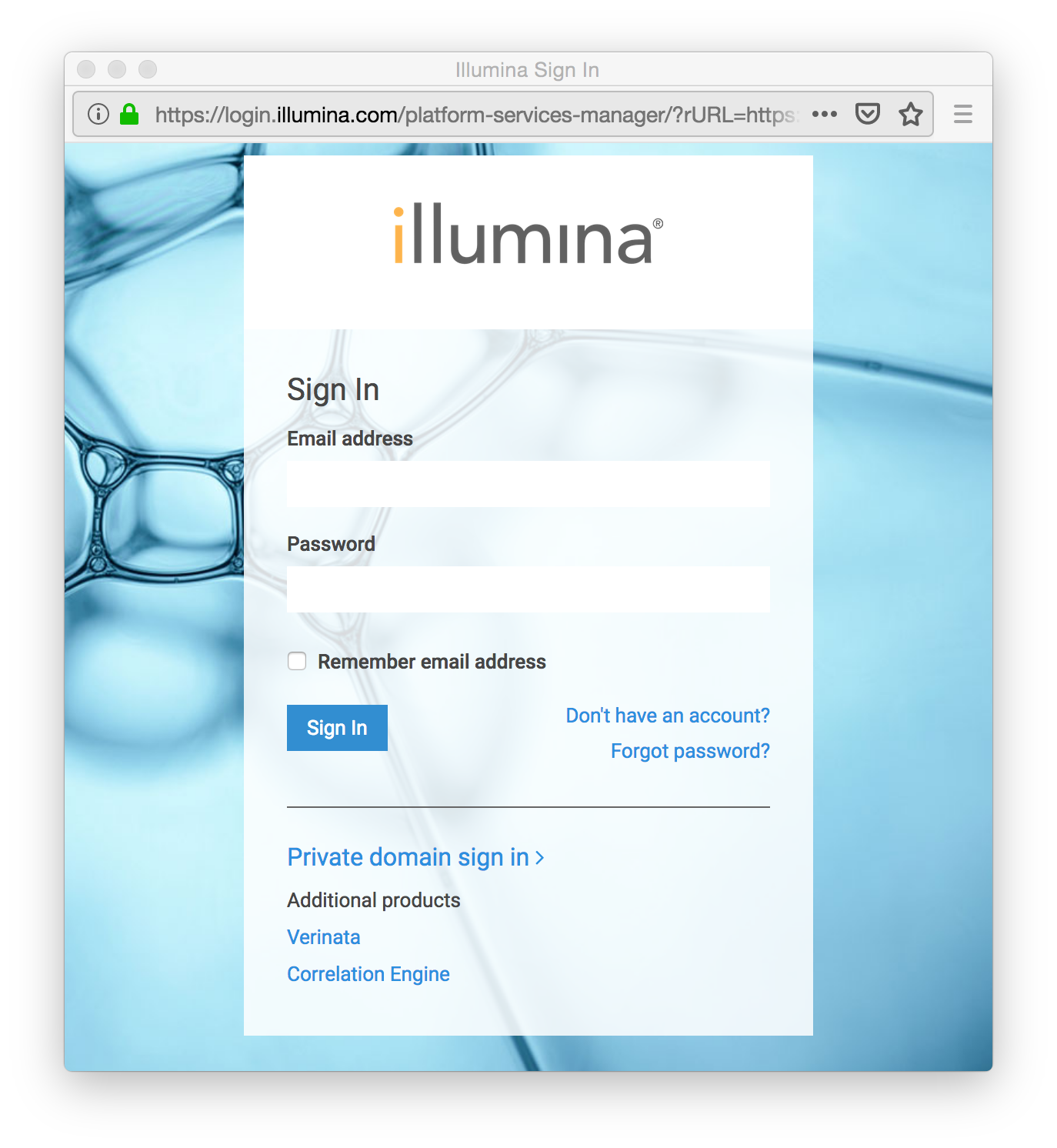
3. Select Samples to Import
Once you have logged in you will see the folders and samples in your BaseSpace account. Click on a folder to view the samples inside. Click on a sample to select it or select multiple contiguous samples by holding down the shift key. You also may select multiple non-contiguous samples by holding down the command (Mac) or Control (PC) key. After selecting the samples you wish to import to Cosmos-Hub, click the “import” button in the upper right corner. After you click “Import” you will be prompted to select your sample type so the proper analysis is performed.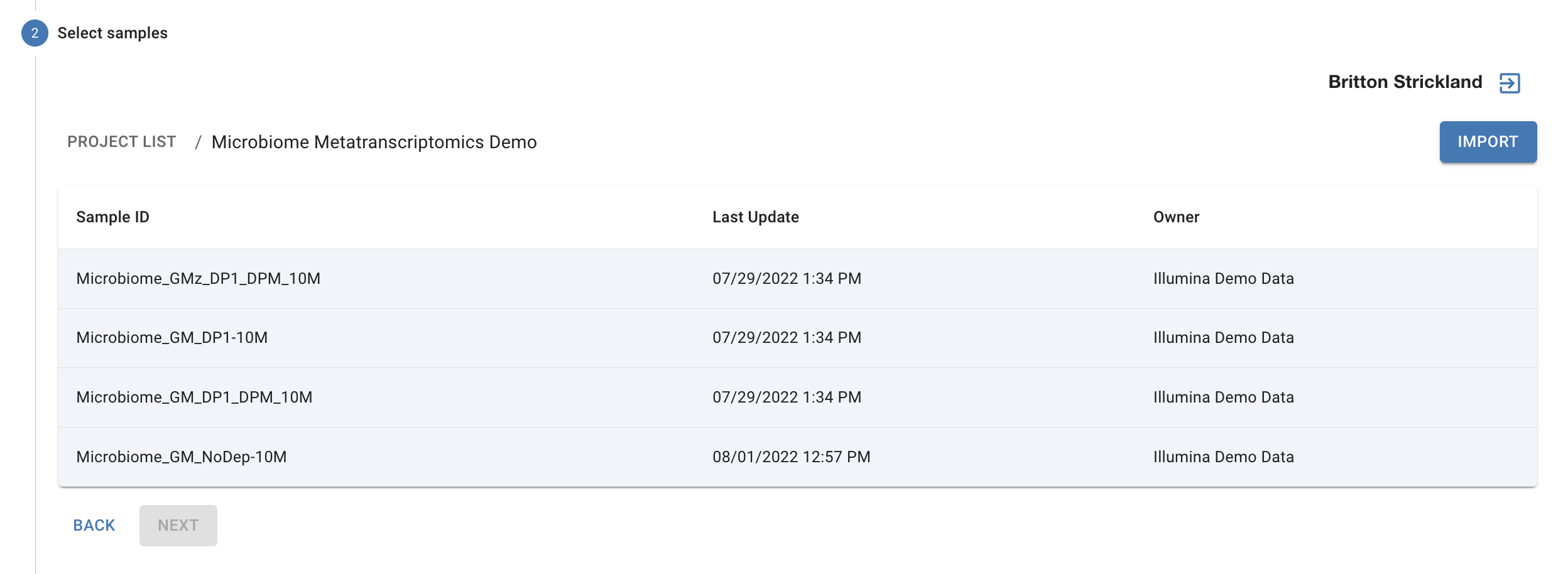
3. Select your desired folder location and data type
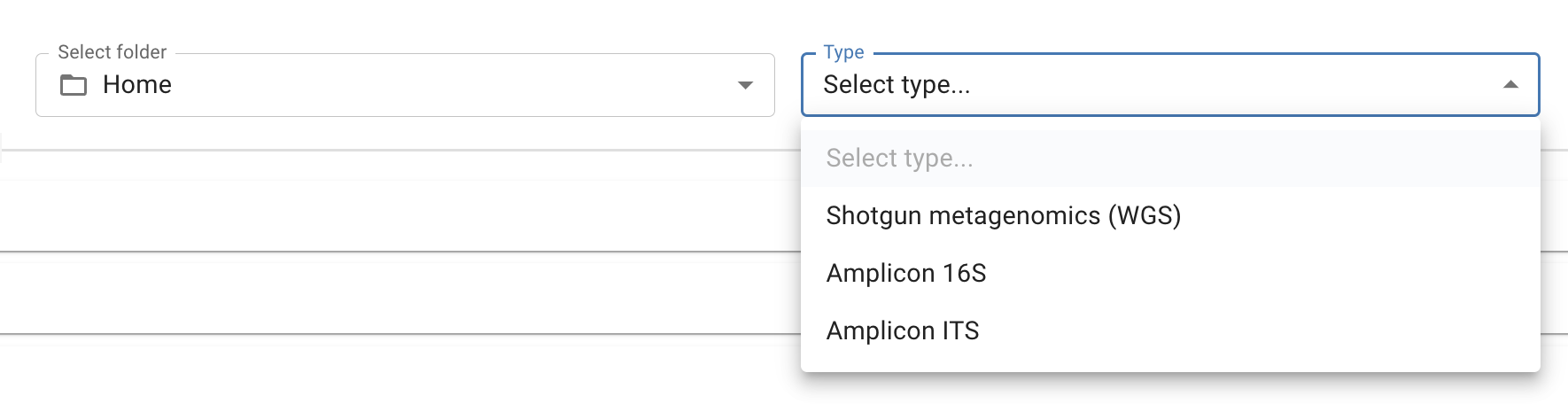
- Shotgun Metagenomics - using whole genome shotgun sequencing, the Cosmos-Hub algorithms identify microorganisms based on entire genomes represented in our database and in your samples
- Amplicon 16S - the amplicon 16S is for bacterial 16S identification. Unlike shotgun metagenomics, amplicon 16S analysis looks only at the relevant bacterial 16S rRNA genes, not the entire genome for identification. This option requires the primer sequences used for generating libraries (e.g,. V1V2, V3V4, V4).
- You can select from standard primer sequences or input your own primer sequence
- Amplicon ITS - the amplicon ITS database is for fungal ITS identification.
3. Remove host reads
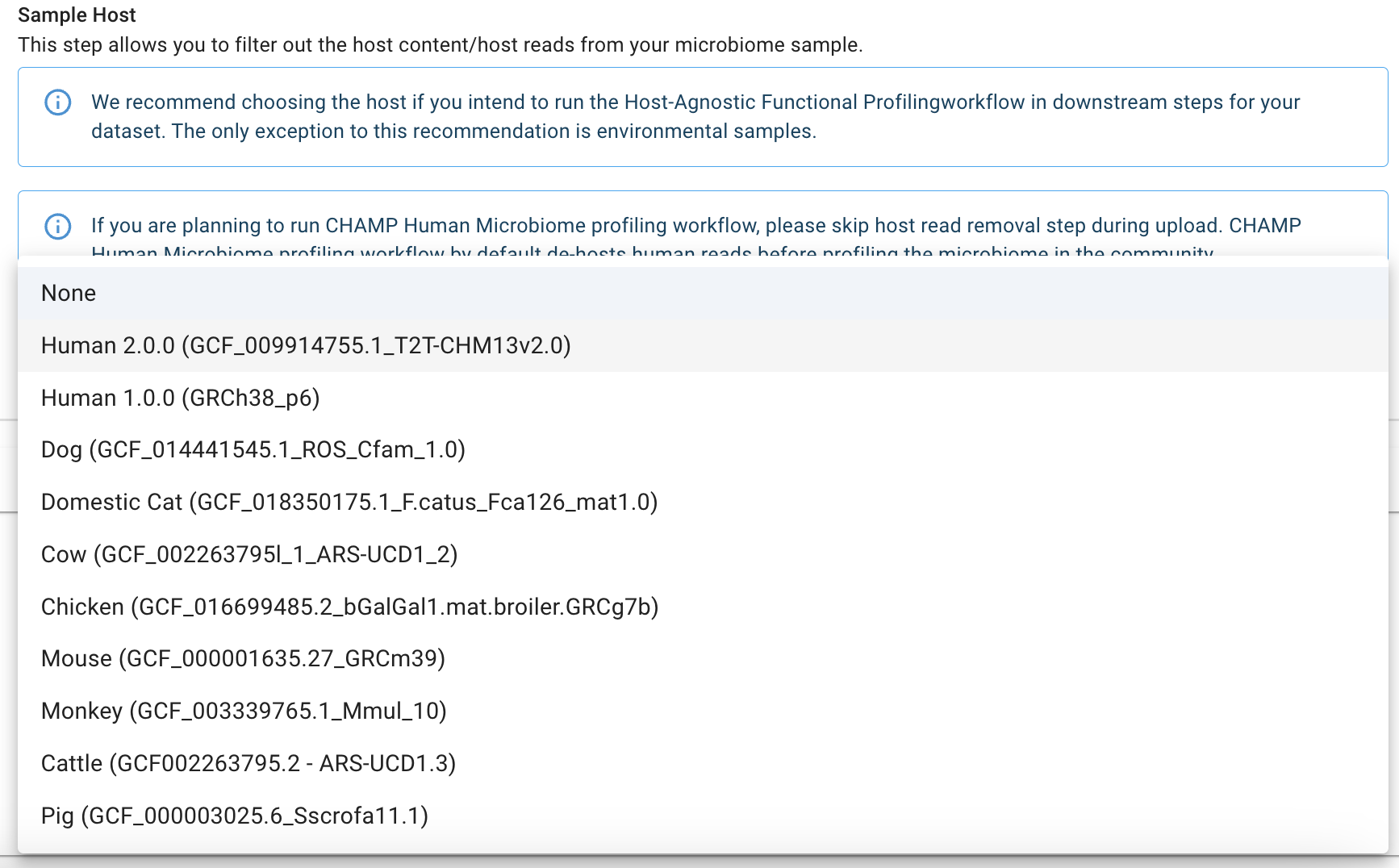
If running CHAMP Human Profiler:
Select “None” as the host when running the CHAMP Human Microbiome Profiler. CHAMP has a native host-removal algorithm for removing human host reads, and selecting a host for this pipeline will result in the sample analysis failing.4. Upload your Metadata (Optional)
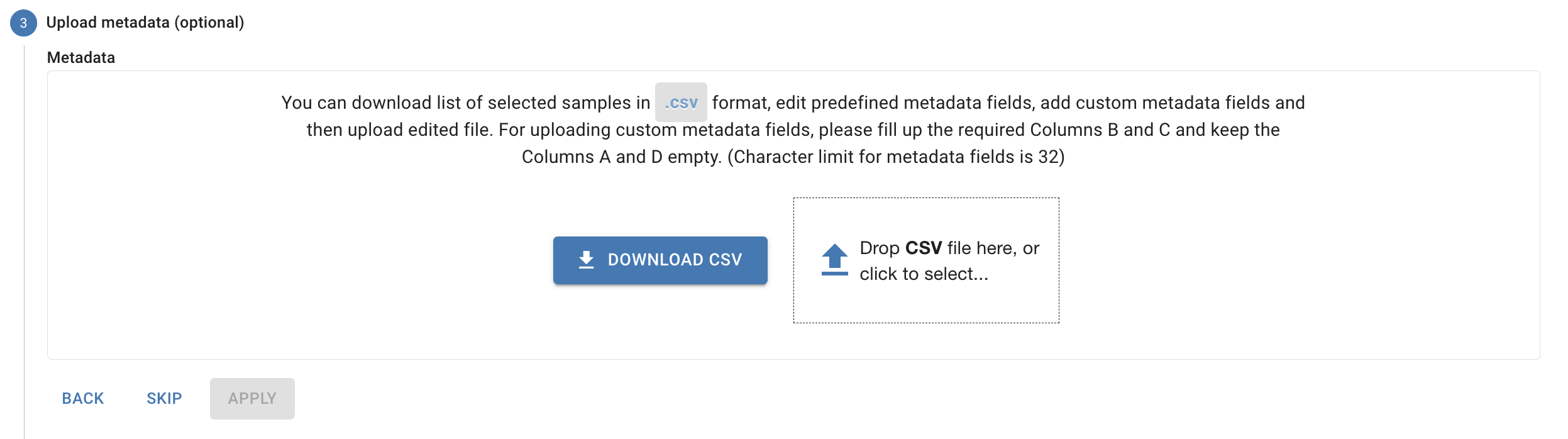
5. Select your workflow
The HUB has several workflows to chose from when uploading your samples:Shotgun Metagenomic Data

Amplicon Data
ASV 16S rRNA Amplicon Profiling OTU ITS rRNA Amplicon Profiling Credit charges are assessed upon successful upload and analysis based on the data type. The Cosmos-Hub team will assign you with the correct number of credits according to your subscription terms. See the chart below for the amount of credits for different analyses.| Taxa | Function | AMR and Virulence Marker | |
|---|---|---|---|
| WGS Deep | 4 | 4 | 2 |
| Amplicon 16S | 1 | N/A | N/A |
| Amplicon ITS | 1 | N/A | N/A |