Why use PCA?
You use PCA to help identify samples that cluster together and form groups based on their composition. For example, if you have many microbiome samples from different body sites and you want to see if they have similar overall bacterial species you can use PCA to determine if they group together based on the body site of origin.How does it work?
PCA simplifies complexity among samples while retaining trends and patterns by transforming the data into fewer dimensions. It finds patterns without prior knowledge about where the samples come from or different variables associated with the samples. PCA projects the data geometrically onto lower dimensions call principal components (PCs). You would typically use PCA plots to find potential clusters and can be used to explore data to understand the key variables in the data and to spot outliers.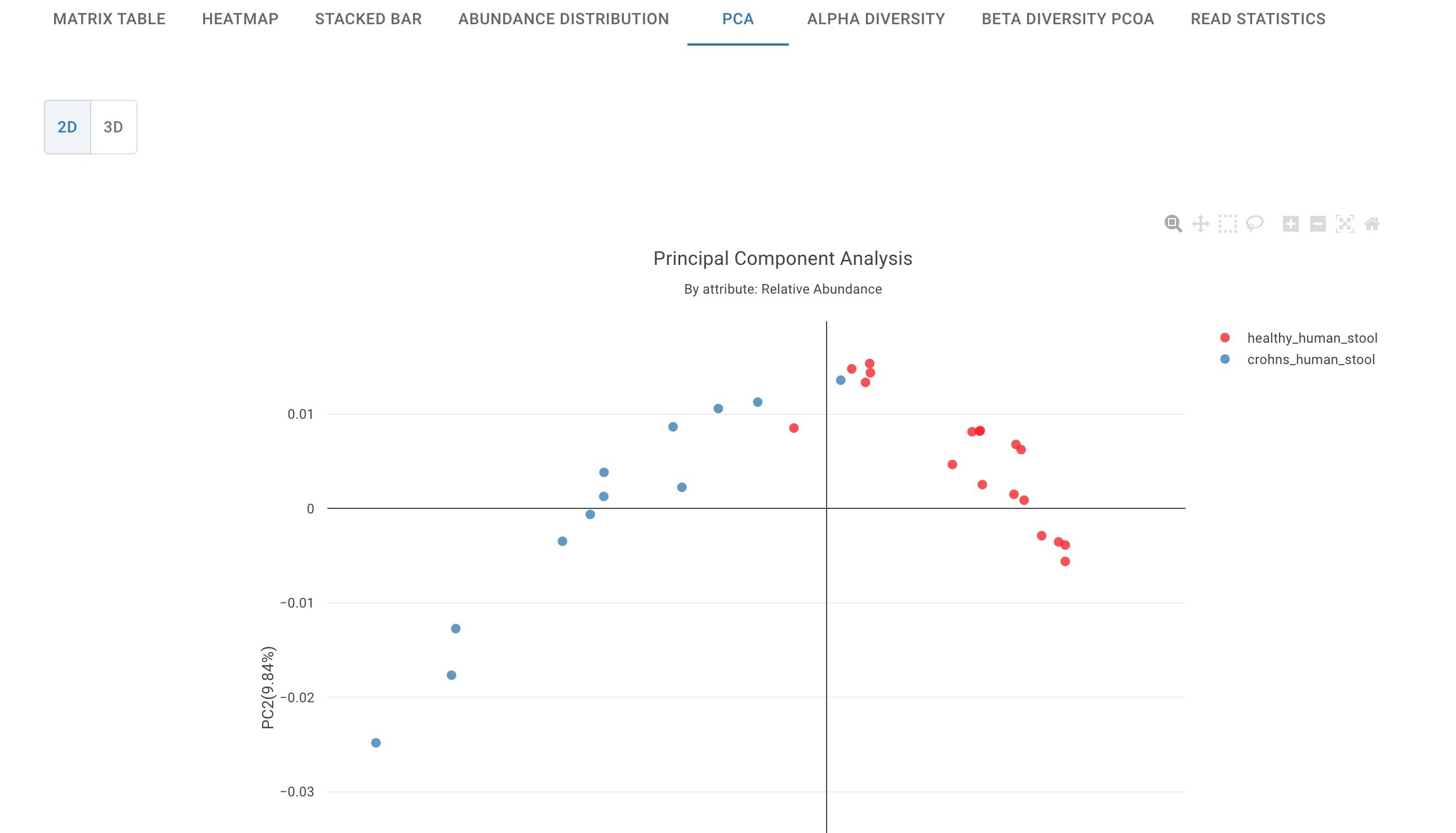
Options for viewing PCA
- Rotate graph - Click anywhere on the 3D PCA and hold down to rotate the graph.
- Labels - You can click on a label to hide samples belonging to the corresponding cohort, click it again to show them.
- Please note that selecting cohort labels in the Legend does not recompute the plot but only hides/reveals the corresponding samples while re-scaling the axis. The axis label values for % variability explained by each principal component (PC) axis are not recalculated and correspond to a visualization including all cohorts and samples.
- Export - Click “Export” in the top right corner to download the PCA as a PDF, PNG or SVG.