Documentation Index
Fetch the complete documentation index at: https://docs.cosmosid.com/llms.txt
Use this file to discover all available pages before exploring further.
You will see the results listed for the database you have selected and you can navigate through all of the databases and sample visualizations such as the sunburst, tree, and stacked bar graphs. Tables vary according to pipeline.
CHAMP Results
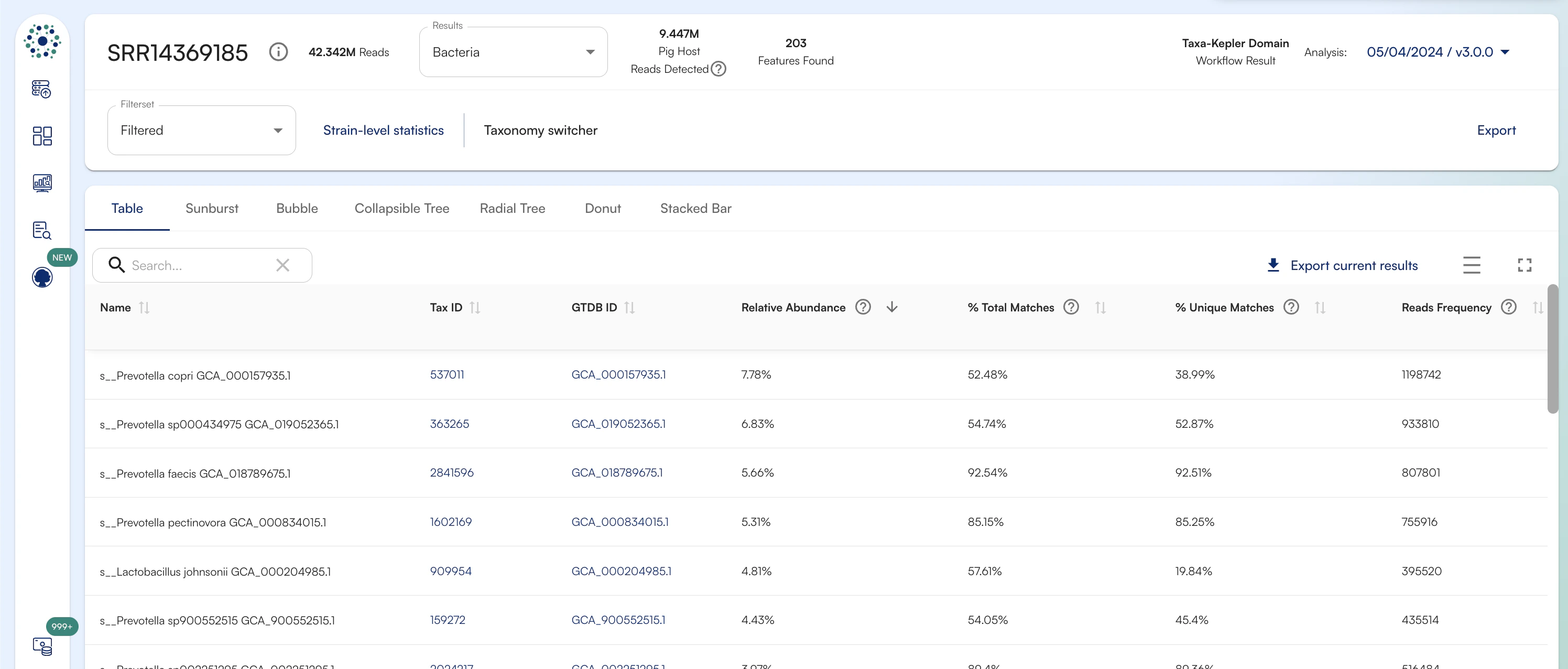
Kepler Results
Definition of Columns
- Name - the name of the organism or functional group, depending on the selected database (taxonomic names based on GTDB nomenclature)
- Relative Abundance - describes the contribution/proportion of a given taxon/feature to the total microbial community detected.
- Counts - number of normalized reads aligning to the reference genome/entry
- Cellular Abundance (functional data) - represents the relative percentage of species that are able to perform a given function in the microbiome community
Definition of Columns
- Name - the name of the organism or taxonomic level or the name of the antibiotic resistance gene or virulence factor, depending on the selected database (taxonomic names based on GTDB nomenclature)
- Tax ID - a link to the NCBI taxonomic ID for the organism
- GTDB ID - a link to the GTDB taxonomic entry for the organism
- Relative Abundance - The Relative Abundance describes the contribution of a given taxon to the total microbial community detected. Relative Abundance is expressed in % and calculated as follows: Relative Abundance = Normalized Reads Frequency (taxon n) / Sum of Abundance Score (all taxa). This metric is suitable for downstream comparative analysis.
- Reads Frequency - number of raw reads aligning to the reference genome/entry
- Normalized Reads Frequency - used to calculate the Relative Abundance (%). This is the number of reads but normalized according to the size of the reference genome. This ensures differences in read counts are not due to larger/smaller genomes and is suitable for downstream comparative analysis or differential abundance analysis.
% Total vs. Unique Matches% Total Matches is the number of shared+unique kmer matches identified in the sample divided by the total number of pre-calculated shared+unique matches available in the reference database. Shared kmers are shared with similar organisms of the same lineage in the phylogenetic tree, which makes this a useful metric for approximating gene coverage, or for assessing the likelihood of a detected taxon or gene actually being present.% Unique Matches is the number of unique kmers found in the sample that are unique to the organism divided by the total number of pre-calculated unique kmers available for the organism in the database. If unique coverage is very low for an organism, it could mean a couple of things:
- For strains, this could indicate that the exact strain in our reference database is not a perfect match to the strain in the sample. However, if you also see a high percent TOTAL coverage for this same strain, this is a good indication that a near taxonomic neighbor of the reference strain is in your sample.
- For some organisms that have high representation in sequencing space (E. coli, for example), there may not be many unique areas in the genome available for strain level identification. As more and more similar genomes are added to the database, we trade-off the ability to discriminate between those that are highly similar to each other. This will cause the percent unique coverage to be low.
This metric is not recommended for downstream comparative analysis.This metric is used by the Filtered setting to rule out likely false positive calls.A common use of % Total Matches is for comparative analysis of antibiotic resistance genes or virulence factors, as these databases are gene-based rather than organism-based. The number of total k-mers identified enables a meaningful in understanding how well the gene has been covered by the reads in the sample.Use the % Total Matches metric to compare how coverage changes between samples, and use the Relative Abundance metric to compare how the composition of marker genes changes between samples.
Other Table Features
- Sort - to sort samples by different columns, click on the column name.
- Search - to search for a sample name, click the magnifying glass and enter your search term.
To understand the top navigation menu and switching taxonomic levels, just go here: View Results.